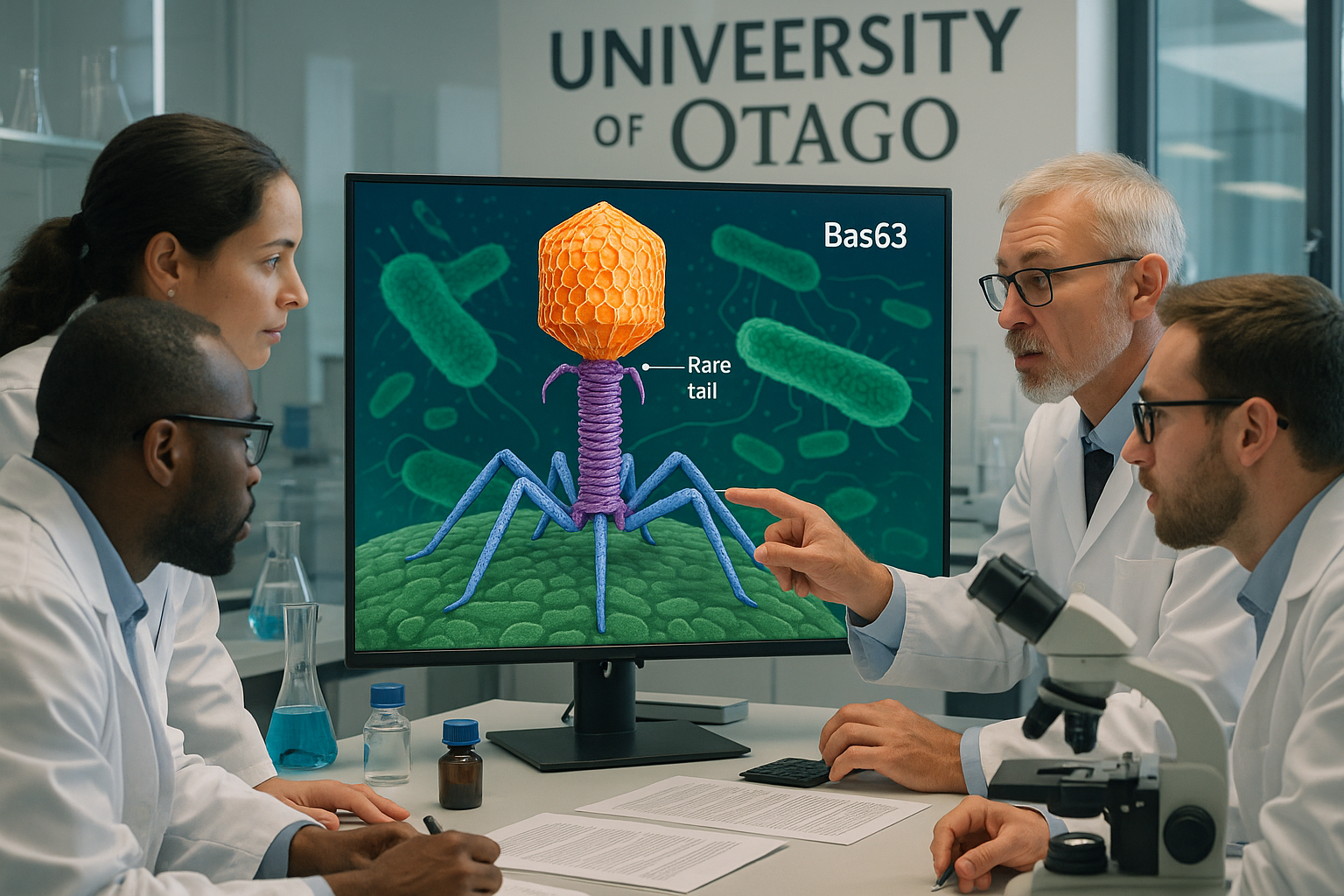
A University of Otago–led team, with collaborators at the Okinawa Institute of Science and Technology, has resolved the 3D structure of Bas63, a bacteriophage that infects E. coli. Published in Science Advances (online on November 12, 2025; issue dated November 14, 2025), the work details rare tail features and could inform rational phage design for medical, agricultural, and industrial uses.
Researchers have produced an in‑depth structural map of the Escherichia phage JohannRWettstein (Bas63), illuminating how its tail apparatus engages bacteria and how related viruses may have evolved. The study appears in Science Advances (DOI: 10.1126/sciadv.adx0790) and lists authors from the University of Otago and the Okinawa Institute of Science and Technology (OIST). Publication records show an online release on November 12, 2025 and an issue date of November 14, 2025. (pubmed.ncbi.nlm.nih.gov)
Lead author Dr. James Hodgkinson‑Bean described bacteriophages as “extremely exciting” alternatives to antibiotics, noting that “bacteriophage viruses are non‑harmful to all multi‑cellular life and able to very selectively target and kill a target bacterium,” which is why they are increasingly studied for phage therapy against drug‑resistant infections. Those remarks were published by the University of Otago and carried by ScienceDaily. (otago.ac.nz)
Using cryo‑electron microscopy, the team reconstructed Bas63 at molecular scale, identifying rare whisker‑and‑collar connections between the head and tail, decoration proteins at hexamer centers on the capsid, and multiple, diverse tail fibers—features that help explain how the virus recognizes and attacks its bacterial host. The OIST news release and the paper’s abstract highlight these elements, including β‑tulip and Hoc‑like decoration proteins, as well as long tail fibers resembling those in phage T4. (oist.jp)
Senior author Associate Professor Mihnea Bostina said the “detailed blueprint of a bacteriophage” can advance rational design for applications ranging from treating infections to combating biofilms in food processing and water systems, a comment echoed in Otago’s newsroom announcement. The researchers also frame the work amid rising antibiotic resistance and threats to global food security from plant pathogens. (otago.ac.nz)
The authors further argue that structural comparisons reveal distant evolutionary links, including relationships between bacteriophages and herpes viruses that may trace back billions of years—a perspective Hodgkinson‑Bean articulated in the university release. The underlying journal article focuses on structural conservation within the Felixounavirus genus and does not date those links, so the time‑scale characterization is presented here as the researchers’ interpretation. (otago.ac.nz)
This is the group’s second full phage structure in 2025: in April, members of the same Otago–OIST team reported the atomic‑level architecture of the potato‑pathogen‑targeting phage φTE in Nature Communications, work they say provides a template for designing biocontrol agents in agriculture. (pmc.ncbi.nlm.nih.gov)